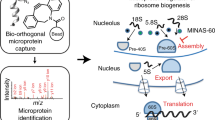
Abstract
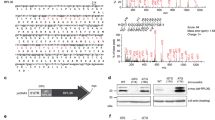
Proteomic detection of non-annotated microproteins indicates the translation of hundreds of small open reading frames (smORFs) in human cells, but whether these microproteins are functional or not is unknown. Here, we report the discovery and characterization of a 7-kDa human microprotein we named non-annotated P-body dissociating polypeptide (NoBody). NoBody interacts with mRNA decapping proteins, which remove the 5′ cap from mRNAs to promote 5′-to-3′ decay. Decapping proteins participate in mRNA turnover and nonsense-mediated decay (NMD). NoBody localizes to mRNA-decay-associated RNA–protein granules called P-bodies. Modulation of NoBody levels reveals that its abundance is anticorrelated with cellular P-body numbers and alters the steady-state levels of a cellular NMD substrate. These results implicate NoBody as a novel component of the mRNA decapping complex and demonstrate potential functionality of a newly discovered microprotein.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Ingolia, N.T., Ghaemmaghami, S., Newman, J.R. & Weissman, J.S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009).
Ingolia, N.T., Lareau, L.F. & Weissman, J.S. Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 147, 789–802 (2011).
Slavoff, S.A. et al. Peptidomic discovery of short open reading frame-encoded peptides in human cells. Nat. Chem. Biol. 9, 59–64 (2013).
Vanderperre, B. et al. Direct detection of alternative open reading frames translation products in human significantly expands the proteome. PLoS One 8, e70698 (2013).
Saghatelian, A. & Couso, J.P. Discovery and characterization of smORF-encoded bioactive polypeptides. Nat. Chem. Biol. 11, 909–916 (2015).
Storz, G., Wolf, Y.I. & Ramamurthi, K.S. Small proteins can no longer be ignored. Annu. Rev. Biochem. 83, 753–777 (2014).
Magny, E.G. et al. Conserved regulation of cardiac calcium uptake by peptides encoded in small open reading frames. Science 341, 1116–1120 (2013).
Anderson, D.M. et al. A micropeptide encoded by a putative long noncoding RNA regulates muscle performance. Cell 160, 595–606 (2015).
Lee, C. et al. The mitochondrial-derived peptide MOTS-c promotes metabolic homeostasis and reduces obesity and insulin resistance. Cell Metab. 21, 443–454 (2015).
Slavoff, S.A., Heo, J., Budnik, B.A., Hanakahi, L.A. & Saghatelian, A. A human short open reading frame (sORF)-encoded polypeptide that stimulates DNA end joining. J. Biol. Chem. 289, 10950–10957 (2014).
Carvunis, A.R. et al. Proto-genes and de novo gene birth. Nature 487, 370–374 (2012).
Fillman, C. & Lykke-Andersen, J. RNA decapping inside and outside of processing bodies. Curr. Opin. Cell Biol. 17, 326–331 (2005).
Sheth, U. & Parker, R. Decapping and decay of messenger RNA occur in cytoplasmic processing bodies. Science 300, 805–808 (2003).
dos Santos, G. et al. FlyBase: introduction of the Drosophila melanogaster Release 6 reference genome assembly and large-scale migration of genome annotations. Nucleic Acids Res. 43, D690–D697 (2015).
Howe, K. et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature 496, 498–503 (2013).
Kent, W.J. BLAT--the BLAST-like alignment tool. Genome res. 12, 656–664 (2002).
Kent, W.J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Thierry-Mieg, D. & Thierry-Mieg, J. AceView: a comprehensive cDNA-supported gene and transcripts annotation. Genome biol. 7 (Suppl. 1), S12.1–S12.14 (2006).
Peng, X. et al. Tissue-specific transcriptome sequencing analysis expands the non-human primate reference transcriptome resource (NHPRTR). Nucleic Acids Res. 43, D737–D742 (2015).
Mellacheruvu, D. et al. The CRAPome: a contaminant repository for affinity purification-mass spectrometry data. Nat. Methods 10, 730–736 (2013).
Parker, R. & Sheth, U. P bodies and the control of mRNA translation and degradation. Mol. Cell 25, 635–646 (2007).
Eulalio, A., Behm-Ansmant, I. & Izaurralde, E. P bodies: at the crossroads of post-transcriptional pathways. Nat. Rev. Mol. Cell Biol. 8, 9–22 (2007).
Fenger-Grøn, M., Fillman, C., Norrild, B. & Lykke-Andersen, J. Multiple processing body factors and the ARE binding protein TTP activate mRNA decapping. Mol. Cell 20, 905–915 (2005).
Eulalio, A., Behm-Ansmant, I., Schweizer, D. & Izaurralde, E. P-body formation is a consequence, not the cause, of RNA-mediated gene silencing. Mol. Cell. Biol. 27, 3970–3981 (2007).
Aizer, A. et al. The dynamics of mammalian P body transport, assembly, and disassembly in vivo. Mol. Biol. Cell 19, 4154–4166 (2008).
Conti, E. & Izaurralde, E. Nonsense-mediated mRNA decay: molecular insights and mechanistic variations across species. Curr. Opin. Cell Biol. 17, 316–325 (2005).
Popp, M.W. & Maquat, L.E. Organizing principles of mammalian nonsense-mediated mRNA decay. Annu. Rev. Genet. 47, 139–165 (2013).
Lejeune, F., Li, X. & Maquat, L.E. Nonsense-mediated mRNA decay in mammalian cells involves decapping, deadenylating, and exonucleolytic activities. Mol. Cell 12, 675–687 (2003).
Li, Y., Song, M. & Kiledjian, M. Differential utilization of decapping enzymes in mammalian mRNA decay pathways. RNA 17, 419–428 (2011).
Lykke-Andersen, J. Identification of a human decapping complex associated with hUpf proteins in nonsense-mediated decay. Mol. Cell. Biol. 22, 8114–8121 (2002).
Lehman, T.A. et al. p53 mutations, ras mutations, and p53-heat shock 70 protein complexes in human lung carcinoma cell lines. Cancer Res. 51, 4090–4096 (1991).
Yamashita, A., Ohnishi, T., Kashima, I., Taya, Y. & Ohno, S. Human SMG-1, a novel phosphatidylinositol 3-kinase-related protein kinase, associates with components of the mRNA surveillance complex and is involved in the regulation of nonsense-mediated mRNA decay. Genes Dev. 15, 2215–2228 (2001).
Martin, L. et al. Identification and characterization of small molecules that inhibit nonsense-mediated RNA decay and suppress nonsense p53 mutations. Cancer Res. 74, 3104–3113 (2014).
Nickless, A. et al. Intracellular calcium regulates nonsense-mediated mRNA decay. Nat. Med. 20, 961–966 (2014).
Brumbaugh, K.M. et al. The mRNA surveillance protein hSMG-1 functions in genotoxic stress response pathways in mammalian cells. Mol. Cell 14, 585–598 (2004).
Keeling, K.M. et al. Attenuation of nonsense-mediated mRNA decay enhances in vivo nonsense suppression. PLoS One 8, e60478 (2013).
Franks, T.M. & Lykke-Andersen, J. The control of mRNA decapping and P-body formation. Mol. Cell 32, 605–615 (2008).
Braun, J.E. et al. A direct interaction between DCP1 and XRN1 couples mRNA decapping to 5′ exonucleolytic degradation. Nat. Struct. Mol. Biol. 19, 1324–1331 (2012).
Ma, J. et al. Discovery of human sORF-encoded polypeptides (SEPs) in cell lines and tissue. J. Proteome Res. 13, 1757–1765 (2014).
Johnson, M. et al. NCBI BLAST: a better web interface. Nucleic Acids Res. 36, W5–W9 (2008).
Karolchik, D. et al. The UCSC Genome Browser database: 2014 update. Nucleic Acids Res. 42, D764–D770 (2014).
Tiscornia, G., Singer, O. & Verma, I.M. Production and purification of lentiviral vectors. Nat. Protoc. 1, 241–245 (2006).
Acknowledgements
We thank A. Ting for the generous use of her Zeiss Axio Observer inverted confocal microscope. This study was supported by a George E. Hewitt Foundation for medical research Postdoctoral Fellowship (Q.C.), NIH (R01 GM102491, A.S.), the NCI Cancer Center Support Grant P30 (CA014195 MASS core, A.S.), The Leona M. and Harry B. Helmsley Charitable Trust grant (#2012-PG-MED002, A.S.), and Dr. Frederick Paulsen Chair/Ferring Pharmaceuticals (A.S.). S.A.S. was supported by Yale University West Campus start-up funds, an American Cancer Society Institutional Research Grant Individual Award for New Investigators from the Yale Cancer Center (IRG-58-012-57) and a NIH Ruth L. Kirchstein postdoctoral fellowship (1F32GM099408). N.G.D. was supported by an Anderson Endowed Postdoctoral Fellowship in the Biological Sciences.
Author information
Authors and Affiliations
Contributions
S.A.S. and A.S. conceived the project. N.G.D., J.M., L.W., Q.C., K.H.L., S.A.S. and A.S. designed the experiments. S.A.S. conducted the majority of the experiments and data analyses, including the identification of NoBody, the discovery of NoBody-binding partners, cellular imaging, and NMD studies. N.G.D. performed NoBody purification and in vitro EDC4 co-purification and crosslinking. J.M. generated NoBody expression constructs, and J.M. and Q.C. carried out the cellular EDC4-NoBody crosslinking experiments. L.W. assisted with identification of NoBody binding partners and co-immunoprecipitation experiments. K.H.L. carried out cellular imaging. E.O.C. and J.L.-A. provided advice and experimental help with NMD studies. B.A.B. provided proteomics assistance. N.G.D., A.S. and S.A.S. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1 and 2, Supplementary Figures 1–16 and Supplementary Note. (PDF 2619 kb)
Supplementary Dataset 1
Immunoprecipitation mass spectrometry data for NoBody pulldowns with anti-FLAG antibody. (XLSX 360 kb)
Rights and permissions
About this article
Cite this article
D'Lima, N., Ma, J., Winkler, L. et al. A human microprotein that interacts with the mRNA decapping complex. Nat Chem Biol 13, 174–180 (2017). https://doi.org/10.1038/nchembio.2249
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2249
This article is cited by
-
Micropeptides: potential treatment strategies for cancer
Cancer Cell International (2024)
-
Small open reading frames: a comparative genetics approach to validation
BMC Genomics (2023)
-
Hydrogel armed with Bmp2 mRNA-enriched exosomes enhances bone regeneration
Journal of Nanobiotechnology (2023)
-
p53-regulated lncRNAs in cancers: from proliferation and metastasis to therapy
Cancer Gene Therapy (2023)
-
A hidden translatome in tumors—the coding lncRNAs
Science China Life Sciences (2023)