Our Services
-
-
-
-
-
-
- Fully customized cell lines in which you specify as many as three proteins of interest (POIs) for tagging
- "Backbone" cell lines where one or two cellular or subcellular structures are pre-tagged and you specify a second or third POI for custom tagging.
- Catalog cell lines in which from one to three POIs are already tagged. View catalog cell lines here.
-
-
-
-
-
Benefits
-
-
-
-
-
-
-
-
- Cell lines delivered 2-3 months faster than competing knock-in technologies
- Up to 3 genes inserted in the same cell
- Your choice of tagging, overexpression models, and point mutations. We can mix the types of knock-ins in the same cell.
-
-
-
-
-
-
-
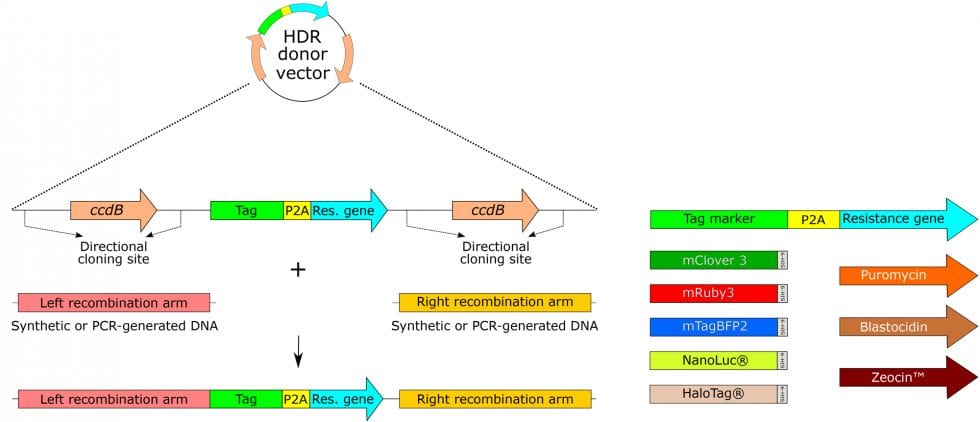
The FAST-HDR Plasmid Vector System:
Although CRISPR/Cas9 has been available for some time now, knocking-in foreign DNA and isolating successfully modified clones remains cumbersome and time-consuming for two main reasons:
-
-
-
-
-
-
-
-
- Creating components used for modifications is time-consuming and requires a high level of expertise.
- Eliminating cells that weren’t modified is necessary to obtain a pure population. This also takes a lot of time.
-
-
-
-
-
-
-
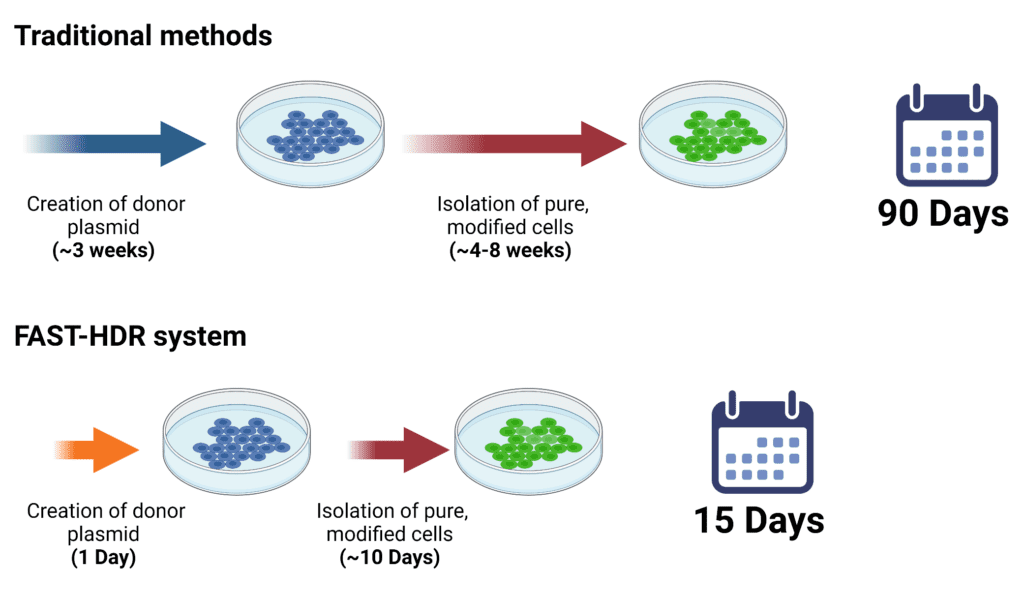
Traditional methods can take 4 – 8 months to deliver a cell line with a single knock-in, twice that long for a cell line with two, and well over a year for a cell line with three knock-ins. Using our FAST-HDR (homology-directed repair) vector system, we can deliver a knock-in cell line 2-3 months faster than conventional methods.
Custom Cell Line Development Service Form

Let's get started
Complete our service request form and we will contact you to discuss your requirements.
FAQs
In the FAST-HDR System, we included a flexible linker sequence that separates the targeted protein from the labeling tag. This approach allows the proteins to be connected and to have spacing to facilitate getting into their correct 3D structure. This approach has worked on labeling essential proteins. However, this approach only partially guarantees that there will be no interference, and sometimes it is required to consider tagging the target protein at the N-term instead of the C-term.
We have done genetic modifications in multiple cancer cell lines that are adherent or grow in suspension. The preferred cell lines have doubling times <30 hours and are a good host to incorporate plasmid DNA with either transfection or electroporation. In addition to working with human cell lines, we have also worked with mouse, rat, and hamster cell lines.
We can use a couple of strategies to generate cell lines with tagged endogenous proteins with low expression. Suppose the expression of the protein can be induced with a small molecule or cytokine. In that case, we can use those molecules to increase expression and facilitate selection during the antibiotic selection step. Alternatively, we can use lower doses of the resistant antibiotics to find a dose that will allow the selection of modified cells with low expression levels of antibiotic-resistant proteins.
- 1 clone of KI cells (2 vials: ≈ 5 x 105cells each)
- 1 vial WT cells (≈ 5 x 105cells)
- Milestone reports sent periodically
- Project report
- Milestone-based project report
- Sequence of guide used
- Primer sequences for PCR and sequencing
- Certificate of Analysis of cells
- Mycoplasma (Pass/Fail)
- Genotype
Luciferase, mClover3, mRuby3, and mTagBFP2 are standard. Other tags are available by special order.
We work with a number of cell lines. Currently, our catalog and backbone offerings are based on HEK293T, U-2 OS, and HeLa cell lines. Examples of other cell lines that we have used include Expi293, HEL92.1.7, OCI-AML3, SNU-899, CHO, and ID8. We have also worked with porcine and bovine cell lines. Please contact us to discuss your cell line options.
Where practicable, we prefer C-terminal tagging since that is less likely to affect endogenous expression of the POI.
Minimal, in that both need the endogenous promoter and the tag needs to be in-frame to be functional.
Absolutely. In our semi-custom cell line service, we stock backbone cell lines with the following combinations of pre-tagged organelles: nucleus and cell membrane; nucleus and cytoskeleton; nucleus and endoplasmic reticulum; and nucleus and mitochondria. You specify the third POI.
It is indeed possible but unlikely to be biologically important since cuts are generally made at the very end of the protein sequence, and the protein will be fully formed until that point.
We use a plasmid containing a high-fidelity Cas9 with high, on-target specificity/low, off-target activity and an optimized sgRNA that must fully interact with the target sequence before cleavage by Cas9. The plasmid, modified to facilitate the cloning of sgRNA oligos, is produced in a single-step reaction with 100% efficiency.
Yes, in those cases in which the function of an essential protein would not be adversely affected by the fluorescent tag. This will only affect timelines modestly since transfection with two donor vectors bearing different resistance genes is done simultaneously.
First, we analyze the sequence of the target gene or genes to determine whether it is biologically possible to effect the cut and the insertion where desired and whether it will be possible to synthesize appropriate recombination arms.
Expression of the antibiotic resistance gene cannot occur without integration. Thus, cells that survive positive selection have been successfully modified.
For HEK293T and HeLa, expression levels for most proteins are known. In certain cases, inducers can be used during clonal selection.
That depends both on the cell line and the POI. Please contact us to discuss.
We use PCR followed by Sanger sequencing.
Western blotting is an optional add-on to the validation workflow.